 Using pROC
Using pROC
Now that pROC is released, it is time to see how it can be used. Here is an example of a typical analysis:
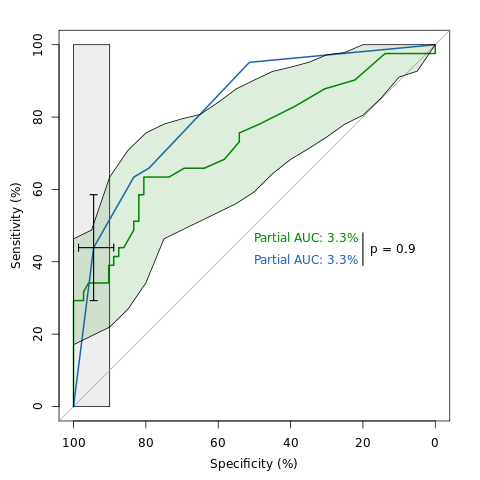 An example of ROC analysis with pROC.
An example of ROC analysis with pROC.
Here is the code that was used to generate this image. The first step is to load pROC (don't forget to type install.packages("pROC") if you haven't installed pROC yet!) and the aSAH data that is provided:
library(pROC) data(aSAH)
First we'll build the ROC curves. We want the sensitivity and specificity in % (thus percent = TRUE) and a partial AUC between 90 and 100% specificity:
rocs100 <- roc(aSAH$outcome, aSAH$s100, percent = TRUE, partial.auc=c(100, 90)) rocwfns <- roc(aSAH$outcome, aSAH$wfns, percent = TRUE, partial.auc=c(100, 90))
Then we compute confidence intervals. For S100, we compute the confidence of sensitivity over several levels of specificity. This will allow us to plot the interval as a shape. With WFNS, we compute the confidence interval of the cut-off 4.5. We use 10000 bootstrap replicates to get a good estimation:
cis100 <- ci(rocs100, of = "se", sp = seq(0, 100, 5), boot.n=10000) ciwfns <- ci(rocwfns, of = "threshold", threshold=4.5, boot.n=10000)
Now, the main feature of pROC is the comparison of two ROC curves. Again we use 10000 bootstrap replicates to get a good estimation:
rtest <- roc.test(rocs100, rocwfns, boot.n=10000)
And finally we plot the ROC curves:
png("demo.png")
plot(rocs100, col="#008600", max.auc.polygon=TRUE, print.auc=TRUE, print.auc.y=48)
plot(rocwfns, col="#1c61b6", add=TRUE, print.auc=TRUE, print.auc.y=42)
And the confidence intervals:
plot(cis100, type="shape", col="#00860022", no.roc=TRUE) plot(ciwfns, type="bars", lwd=1.5)
Last we add the p value of the ROC comparison test:
text(18, 45, sprintf("p = %.1f", rtest$p.value), adj=c(0,1))
lines(c(20, 20), c(39, 48), lwd=1.5)
dev.off()
And that's it, we have generated the image above!
You can find more such examples on the screenshots page on ExPASy!
Xavier Robin
Publié le mardi 27 avril 2010 à 20:50 CEST
Lien permanent : /blog/2010/04/27/using-proc
Tags :
pROC
Commentaires : 0
 pROC 1.0 released
pROC 1.0 released
After years of work with ROC curves in R, first with packages such as ROCR and verification, then with custom functions; and 4 months of development to create a coherent package, extensive discussions about its name (SwissROC was the first thought (but is Switzerland still a selling argument?), or Rocker (that's a bit similar to the already-existing ROCR)) and its licensing (let's go with GPL), I'm glad to announce the release of pROC!
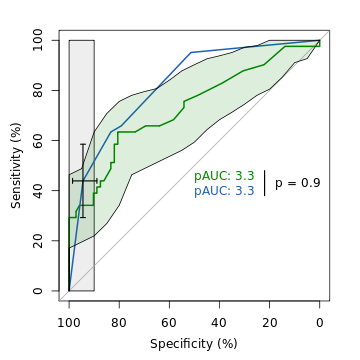 An example of ROC analysis with pROC.
An example of ROC analysis with pROC.
pROC is a set of tools to visualize, smooth and compare receiver operating characteristic (ROC curves). Full or partial area under the curve (AUC) can be compared with statistical tests based on U-statistics or bootstrap. Confidence intervals can be computed for (p)AUC or ROC curves. It is available for R (command-line interface) and S+ (with an additional graphical user interface).
You can find it on ExPASy (thanks Céline from the SIB web-team) and in a few minutes on the CRAN (it has to pass a few checks before it is formally accepted).
But there is no need to download the package. The installation can be done in one command directly from R:
install.packages("pROC")
The package must then be loaded with:
library(pROC)
To get help, enter the following in the R prompt:
?pROC
And that's it! You can have a look at the screenshots to see what it looks like.
Simultaneously we submitted a paper (a short application note) describing it to Bioinformatics. Crossed fingers in the hope it will be accepted!
2010-06-15 follow-up: the paper has been rejected. We will resubmit it to BMC Bioinformatics.
2011-03-17 follow-up: the paper was published in BMC Bioinformatics.
Xavier Robin
Publié le mardi 27 avril 2010 à 20:42 CEST
Lien permanent : /blog/2010/04/27/proc-1.0-released
Tags :
pROC
Commentaires : 0
 Chercher
Chercher Tags
Tags Billets récents
Billets récents Calendrier
Calendrier Syndication
Syndication